Biomechanical modeling is the development of a mathematical representation of the human body and simulation is the process of running experiments on the biomechanical model. There are two main types of modeling used in biomechanical analysis:
- Inverse dynamics – This is the much more widely used and recognized type of biomechanical modeling. Inverse dynamics relies on measuring the motion of a subject in a clinical, orthopedic, or sports movement trial and combining the measured motion data with a body model to calculate (estimate) the forces that were necessary to produce this movement. Standard inverse dynamics, due to its dependence on measured motion data, is not well suited to predict outcomes. The term inverse is a reminder that flow of calculations in this analysis is opposite to the way in which movement is actually produced in the body. Traditional biomechanical methods for inverse dynamic computation for high-speed sports motions often suffer due to inherent limitations of processing motion capture data as well as the complexity of the 3D dynamic equations of motion for non-orthogonal models of anatomical motion that include an anatomically relevant pronation/supination axis. I personally have had success using a new method called Instantaneous Torque Induced Accelerations (ITIA) for these demanding modeling applications.


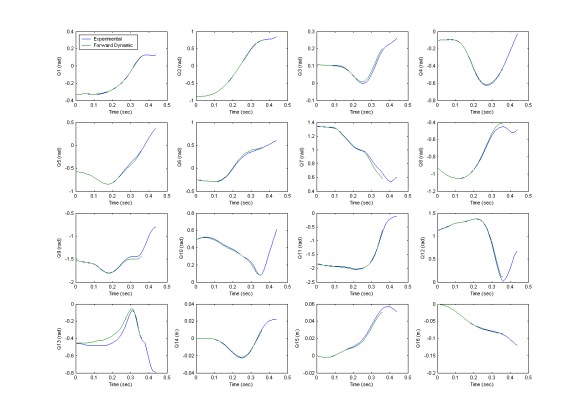
- Forward dynamics – This type of biomechanical modeling is not as widely used, though in reality, it should be as it is a much more powerful modeling technique due to its predictive capabilities. Forward dynamics uses joint torques/forces to predict resultant motions. The reason why this is such a powerful predictive tool is that the software logic mimics the manner in which the human body actually functions. In real life, neural inputs give rise to muscle forces that generate joint and ground reaction forces that together drive the movement of the human body. This order is maintained in the forward dynamics or simulation paradigm. And that is why forward dynamics modeling is such a powerful predictive tool. The Kane’s Method of forward dynamic simulation is powered by the active joint torques calculated by the ITIA method and is controlled by the passive joint torques specific to each subject. As such, the resultant accuracy of the forward dynamic simulation will be completely dependent upon the effectiveness of the inverse dynamic method and the estimation of the passive joint torques. The ITIA method has proven to be very robust for high-speed sports motions, with simulations yielding R-Factors between 0.95 – 1.00 for all degree of freedom (DOF) between the forward dynamic orientations and the original experimental data as shown below.
I have over 10 years of experience in the development and implementation of dynamic musculoskeletal models using the vectorized kinematic approach of Kane’s Method as well as the trigonometric kinematic approach of Newton-Euler and Lagrangian formulations for the dynamic equations of motion. Given a choice, I will always lean to using Kane’s Method as the vectorized approach is much easier to use and implement with modern day computer software programs and is used in exactly the same way to solve every problem. Using Kane’s Method, I am able to spend much more time on meaningful biomechanical analysis as opposed to spending my time on creating and debugging the dynamic equations of motion for difficult 3D problems. Due to the inherent advantages of Kane’s Method, it is much easier to develop more complex 3D biomechanical models which provide ever more realistic representations of the physiological system being modeled.
One of the limitations with Kane’s Method is limitations with animation and visualization capabilities. There are really no turnkey software solutions that provide high-end animation capabilities. It is necessary to use other software packages with animation capabilities if visualization is an important consideration, such as SIMM by MusculoGraphics. The quantitative outputs from Kane’s Method analyses are still what makes this the de facto method of choice for the majority of forward dynamics analyses. However, I have also used LifeMOD with much success for a variety of forward dynamics projects due to a significant advantage in animation and synchronized kinematic and kinetic output visualization capabilities. The trade-off is in control of the inverse dynamics method, where LifeMOD uses a psuedo-inverse methodology vs. a traditional inverse dynamics methodology. I have found that Kane’s Dynamics methodology allows me greater control over the inverse dynamics process which results in quicker production of forward dynamics simulations with very high correlation coefficients between simulation results and input motion data. I have also had success with LifeMOD, however, it takes a little more time and intuition to control the pseudo-inverse method to get similar results. But the animation and visualization capabilities of LifeMOD definitely provide the difference for some application projects. I have used LifeMOD for a number of application projects for sports equipment performance as well as orthopedic device studies.
As many of my forward dynamic simulations are used for sports equipment performance, joint torque driven models are preferred as the analysis is focused on resultant motion of the equipment itself. For the golf swing, a joint torque driven model swings the club exactly the same way every time, allowing parametric analysis of golf club parameters for a variety of swing types. For baseball, a subject-specific model can be used to analyze resultant ball spin for different pitch types for comparison to PITCHf/x data to analyze player specific spin optimization.
However, there are times where muscle actuators are included in the model to replace the active joint torques driving the simulation.. This extends the analysis to studying actual forces required by each muscle to drive the simulation. I typically use these types of simulations when I am interested in micro-level joint detail, such as ligament loads or activation patterns in specific muscle groups. However, it is much more difficult to validate these simulation outputs as it is very hard to get any EMG validation in high-speed sports movements. In active joint torque driven models, one can validate the accuracy of the simulation by comparing simulation outputs to measured data. It is impossible to validate these outputs as we can’t directly measure dynamic muscle loads and timings. Nonetheless, these simulations are very powerful for demonstrating optimal muscular activation patterns for a variety of motion patterns.
I have considerable experience using advanced symbolic manipulator software programs (MotionGenesis Kane, AUTOLEV, and Maple) for biomechanical model derivation and programming of the equations of motion (MATLAB, C/C++) as well as turnkey software solutions (ADAMS, FigMOD, LifeMOD).